•
the Blog
Against recruitment panel interviews
By Guillaume Filion, filed under
recruitment,
management.
• 27 January 2017 •
 In my first years as a group leader, I had the chance to interview PhD candidates in panels at international calls for students. I quickly stopped interviewing students, but back then I was very surprised that the top candidates often proved less productive than those we had ranked mediocre. How was this possible at all? Panels are unbiased, they combine multiple expertises, they allow for critical discussion, so they should be able to pick the best candidates... right?
In my first years as a group leader, I had the chance to interview PhD candidates in panels at international calls for students. I quickly stopped interviewing students, but back then I was very surprised that the top candidates often proved less productive than those we had ranked mediocre. How was this possible at all? Panels are unbiased, they combine multiple expertises, they allow for critical discussion, so they should be able to pick the best candidates... right?
It turns out to be less surprising than I thought. Now a little more familiar with the dangers of panel interviews, I decided to see what our colleagues from the academia have to say about it. This is of course where I should have started before interviewing PhD students, but better late than never. If you haven’t met them, let me introduce you to the flaws of the mighty recruitment panel interview...
1. The unproved efficiency of panels
A good place to start. Despite forty years of research, the benefit of recruitment panels over single evaluators is still debated. According to a review published in 2009
Findings to date suggest that panel interviews might not necessarily provide the psychometric...
Three lessons from peer coaching
By Guillaume Filion, filed under
motivation,
coaching,
leadership,
management.
• 09 January 2017 •
 For a little more than a year, my colleagues and me have been organizing peer coaching sessions for junior group leaders. A typical session consists of four to six of us, and we meet for one morning to discuss the most pressing issues. After a start-up training and some trial and error, we settled for a group coaching method that gave the best result. To give an idea, the “coachee” tells the chairman what he/she wants to solve, then follows a discussion where he/she explains the facts to the coaches who ask as many question as possible. Then the coaches analyze the situation, suggest solutions and make comments meanwhile the coachee has to remain silent and listen. Finally, the coachee summarizes what he/she heard and what steps he/she will take.
For a little more than a year, my colleagues and me have been organizing peer coaching sessions for junior group leaders. A typical session consists of four to six of us, and we meet for one morning to discuss the most pressing issues. After a start-up training and some trial and error, we settled for a group coaching method that gave the best result. To give an idea, the “coachee” tells the chairman what he/she wants to solve, then follows a discussion where he/she explains the facts to the coaches who ask as many question as possible. Then the coaches analyze the situation, suggest solutions and make comments meanwhile the coachee has to remain silent and listen. Finally, the coachee summarizes what he/she heard and what steps he/she will take.
With this exercise, we learned a great deal about how to organize such peer coaching sessions in the academia and how to make the best of them, but this is not what this post is about. Instead, I would like to share more important lessons I have learned about working together and using the group as support and source of motivation.
If you...
One or two tails? 
By Guillaume Filion, filed under
statistics,
p-hacking.
• 11 December 2016 •
 Here is a discussion that I recently had with my colleague John. He approached me with the following request:
Here is a discussion that I recently had with my colleague John. He approached me with the following request:
A tutorial on Burrows-Wheeler indexing methods (1) 


By Guillaume Filion, filed under
full text indexing,
Burrows-Wheeler transform,
suffix array,
series: focus on,
bioinformatics.
• 04 July 2016 •
There are many resources explaining how the Burrows-Wheeler transform works, but so far I have not found anything explaining what makes it so awesome for indexing and why it is so widely used for short read mapping. I figured I would write such a tutorial for those who are not afraid of the detail.
The problem
Say we have a sequencing run with over 100 million reads. After processing, the reads are between 20 and 25 nucleotide long. We would like to know if these sequences are in the human genome, and if so where.
The first idea would be to use grep to find out. On my computer, looking for a 20-mer such as ACGTGTGACGTGATCTGAGC takes about 10 seconds. Nice...
Did Mendel fake his results? 
By Guillaume Filion, filed under
statistics,
fraud,
p-hacking,
genetics.
• 11 April 2016 •
 You went to high school and you learned genetics. You heard about a certain Gregor Mendel who crossed peas and came up with the idea that there is a dominant and a recessive allele. You did not particularly like the guy because there would always be a question about peas with recessive and dominant alleles at the exam. But you grew up, became wiser and just as you started to like him, you heard from someone that he faked his data. You felt disoriented for a while, why annoy you with this stuff at school if it is wrong? But then you came to the conclusion that he just got lucky and that he was right for the wrong reasons. After all, he was just a monk on gardening duties, why would you expect him to understand anything about real science?
You went to high school and you learned genetics. You heard about a certain Gregor Mendel who crossed peas and came up with the idea that there is a dominant and a recessive allele. You did not particularly like the guy because there would always be a question about peas with recessive and dominant alleles at the exam. But you grew up, became wiser and just as you started to like him, you heard from someone that he faked his data. You felt disoriented for a while, why annoy you with this stuff at school if it is wrong? But then you came to the conclusion that he just got lucky and that he was right for the wrong reasons. After all, he was just a monk on gardening duties, why would you expect him to understand anything about real science?
Gregor Mendel
Gregor Mendel was a monk, but he was also a trained scientist. He studied assiduously for twelve years (including about seven years on physics and mathematics), to then become a teacher of physics and natural sciences at the gymnasium of Brno. He prepared his most famous experiment for two years, meticulously checking and choosing his...
How to teach research?
By Guillaume Filion, filed under
motivation,
leadership,
management.
• 28 February 2016 •
 One of my students once asked me how to become a better researcher. “Great question!” I thought. Instead of a straight and clear answer, I heard myself babbling, as happens when you have no idea what you are talking about. Somewhat disappointed by my non answer, I turned to my colleagues and asked how they teach research. The answer ranged from embarrassed silence to politely switching topic. Strange, I thought, it is our responsibility as principal investigators to educate the junior researchers of our group, so we’d better have some idea of what we are doing.
One of my students once asked me how to become a better researcher. “Great question!” I thought. Instead of a straight and clear answer, I heard myself babbling, as happens when you have no idea what you are talking about. Somewhat disappointed by my non answer, I turned to my colleagues and asked how they teach research. The answer ranged from embarrassed silence to politely switching topic. Strange, I thought, it is our responsibility as principal investigators to educate the junior researchers of our group, so we’d better have some idea of what we are doing.
We have a fairly good idea of how to teach skills, like mathematics, molecular biology etc. But I would argue that research is not a skill. I would argue that it is an attitude. So the real question is how to teach an attitude. There is no foolproof recipe, but here goes a subjective view that I gathered from reading, from discussions and from personal experience.
1. Motivate the change
Most students and junior scientists enter the lab with an open mind, ready to learn as many new things as possible. But most of them focus on learning technical skills (at...
ENCODE data, Principal Components and racism
By Guillaume Filion, filed under
racism,
series: genetics and racism,
Principal Component Analysis,
ENCODE.
• 28 September 2015 •
 “Thinking is classifying” wrote Georges Clémenceau*. This tells, in simple words, everything about the obsession of the human mind to keep things tidy. No surprise we ask computers a little help here and there. Is this email spam? Is this online user human? Is this text written by that author? Training machines to put things into the boxes created by our human mind is called supervised learning and it can be very lucrative. But what about the more philosophical cases where machines make their own boxes? Can we reverse the process and put things in boxes created by computers? Unsupervised learning, as it is called, creates a lot of interesting problems where we, humans, are left wondering whether the boxes make any sense.
“Thinking is classifying” wrote Georges Clémenceau*. This tells, in simple words, everything about the obsession of the human mind to keep things tidy. No surprise we ask computers a little help here and there. Is this email spam? Is this online user human? Is this text written by that author? Training machines to put things into the boxes created by our human mind is called supervised learning and it can be very lucrative. But what about the more philosophical cases where machines make their own boxes? Can we reverse the process and put things in boxes created by computers? Unsupervised learning, as it is called, creates a lot of interesting problems where we, humans, are left wondering whether the boxes make any sense.
The mother of all classification techniques is undisputedly Principal Component Analysis (PCA). But let me reassure those who hate PCA and those who never heard of it: I will just touch the surface, and then very briefly. PCA automatically arranges similar items close to each other on a plane. The rest is up to you. Similarity, in particular, depends on a bunch of arbitrary features, size, height, number of legs... In a classical introductory example...
Stick breaking and DNA alignment 

By Guillaume Filion, filed under
spacings,
stick breaking,
heuristic,
sequence alignment,
bioinformatics.
• 06 September 2015 •
A couple of months ago, I posted an approximate formula for the longest match in the problem of DNA alignment. I recently used it to calibrate a seeding heuristic to map Illumina reads and I was surprised to see that it was not just bad, but epic bad. Upon closer inspection, I realized that the main assumption does not hold when the error rate is small (which is typically the case for Illumina reads). The formula was based on longest runs in Bernoulli trials. This time I present more accurate results with an approach based on a stick breaking process.
Stick breaking (spacings)
Inserting $(k)$ mutations at random in a sequencing read will produce $(k+1)$ (possibly empty) subsequences without errors. The process is analogous to inserting $(k)$ breaks at random in a stick of length 1, and we can approximate the distribution of the longest subsequence without error by that of the longest fragment when breaking the stick.

The example above illustrates this concept graphically. A sequencing read of 60 nucleotides contains 2 mutations highlighted in red and the longest error-free stretch is the central subsequence of 28 nucleotides. If mutations occur uniformly on the read...
Bayesian networks and causation 
By Guillaume Filion, filed under
statistics,
correlation,
Bayesian networks,
causes.
• 20 June 2015 •
 The first thing you learn in statistics is that “correlation does not imply causation”. As obvious as it sounds, most human mistakes fall in this category, and not only in statistics. The major difficulty with this question is that it is fairly easy to define correlation, but it is much harder to define causation, let alone quantify it. No surprise many statisticians just avoid talking about causation to stay out of the danger zone.
The first thing you learn in statistics is that “correlation does not imply causation”. As obvious as it sounds, most human mistakes fall in this category, and not only in statistics. The major difficulty with this question is that it is fairly easy to define correlation, but it is much harder to define causation, let alone quantify it. No surprise many statisticians just avoid talking about causation to stay out of the danger zone.
However, for Judea Pearl, this is not a satisfactory answer. In his book Causality: Models, Reasoning, and Inference, he expresses his opinion vividly.
I see no greater impediment to scientific progress than the prevailing practice of focusing all our mathematical resources on probabilistic and statistical inferences while leaving causal considerations to the mercy of intuition and good judgment.
This book lays the foundation of the now popular Bayesian networks. The key idea is that you can distinguish correlation from causation if you can observe several independent causes. For instance, suppose that patients suffering from a certain type of cancer are often immunodeficient. You wonder whether immunodeficiency is a cause or a consequence of this cancer type.
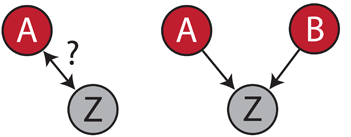
Say that variable A is whether patients have...
What is bioinformatics about?
By Guillaume Filion, filed under
PubMed,
bioinformatics,
journals,
information retrieval.
• 01 June 2015 •
A brief note published a few weeks ago initiated a discussion on the blogosphere about who is a bioinformatician and who is not. According to Wikipedia
bioinformatics combines computer science, statistics, mathematics, and engineering to study and process biological data.
To see how the community defines itself, I downloaded the abstracts of Bioinformatics from 2014 (for a total around 1000 articles, extracted the most relevant keywords and put the top 100 terms in a word cloud where a word size shows its frequency*.

Obviously, bioinformatics is about data, mostly gene/protein sequences and expression. It is also good to know that bioinformatics likes genomes and networks, and that it has more affinities with structural biology than with evolution.
The favourite organism of bioinformaticians is Homo sapiens, actually it is the only one mentioned in the word cloud, and when bioinformaticians work on a disease, it is cancer.
When bioinformaticians describe their work, it is “novel” and “new”, and what they talk about is “biological”, “different”, “multiple” and “single” (the last two are usually followed by “sequence alignment” and “nucleotide polymorphism”). It is also “functional” and “available”, but somewhat less. I expected it to be “fast” and “accurate”, but...

